Semler, E. M., Michell, D. L., Kingsley, P. J., Ramirez, M. A., Massick, C., Castleberry, M., Doran, A. C., Carr, J. J., Marnett, L., Sheng, Q., Linton, M. F., & Vickers, K. C. (2026).ä».ä»Atherosclerosis,ä»412, 120584.ä»
Chemical modifications on RNA, calledô epitranscriptomic modifications, help control how RNA works in our cells. Small RNA fragments, known asô tRNA-derived fragments (tDRs), can carry these modifications from their parent RNA. Some of these modified RNAs travel through the blood insideô high-density lipoproteins (HDL), the same particles that carry ãgood cholesterol.ã These HDL-associated RNAs (HDL-sRNAs) can influenceô immune responsesô and may play a role inô atherosclerotic cardiovascular disease (ASCVD).
In this study, we compared HDL-sRNAs from healthy people and patients with advanced ASCVD. Using methods likeô LC-MS/MS,ä»ARM-seq, andô qPCR, we found that HDL from ASCVD patients had more RNA modifications. We identified a specific fragment,ä»tDR-ArgACG, carrying a modification calledô 1-methyladenosine (m1A), which was more common in ASCVD. When these modified RNAs enteredô macrophages, immune cells that drive inflammation in arteries, they changed gene activity, including increasingô TMEM123, a protein that helps immune cells move and stick to tissues. Experiments showed that HDL carrying m1A-tDR-ArgACG boosted TMEM123 in macrophages, and blocking the m1A modification reduced this effect.
These findings suggest that HDL can deliver modified RNAs to immune cells, triggeringô inflammationô in atherosclerotic plaques. This reveals a new way that HDL-sRNAs may contribute to cardiovascular disease and points to potential targets for therapy.
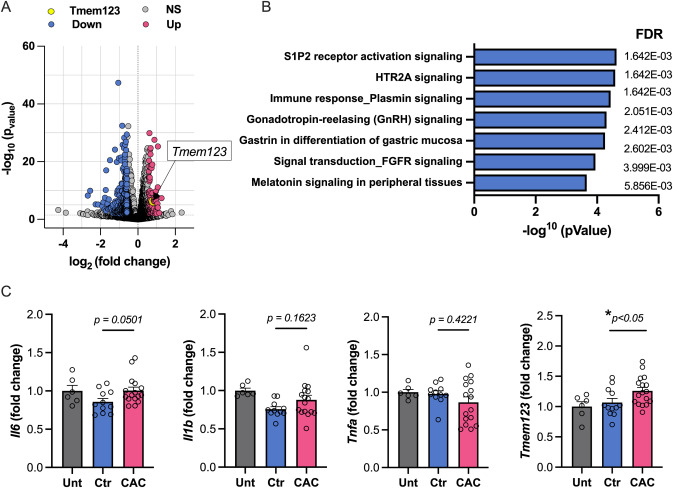
Fig. 1ô CACô HDL induces non-classical activation and select immune response pathway and gene expression changes in macrophagesô (A) Volcano plot depicting log10 p-adjusted (Q)-value versus log2 fold changes for all differentially expressed (ãË2.0 absolute fold change) genes in BMDM’s treated with CACô +ô HDL or Ctr-HDL (nô =ô 3). (B) EGR1-containing gene set pathway analyses (Meta Core) enrichment, ranked by -log10(p-value) with false discovery rate (FDR) shown. (C) mRNA expression by qPCR displaying fold change from BMDMs treated with Ctr-HDL (nô =ô 9) and CAC+HDL (nô =ô 14). Statistical analysis assessed by Mann-Whitneyô Uô test. Data are presented as meanô +s.e.m., ãÝÒä»&ݶ°ì;ä»0.05.